The growth of artificial intelligence (AI) tools and machine learning (ML) models in drug screening has dramatically increased the number of viable drug targets while slashing the time to market. This shift has fueled a boom in dry labs—computational environments where simulations and mathematical analyses replace the pipettes, reagents, and spectrometers in traditional wet labs.
In Canada, this evolution has created a unique geographical challenge. Although Toronto has solidified its status as a global dry lab and AI powerhouse (anchored by the Vector Institute and Roche’s Data Science Coalition), a bottleneck persists. Once a promising candidate is identified, there may be no nearby wet lab space to continue its research and development. This is why every new wet lab space that opens in the region becomes newsworthy. Another bottleneck is that the in silico models generated with AI aren’t always in a format that meshes easily with wet lab analytical results.
For Canadian biopharmaceutical organizations forecasting growth in the next five years, this gulf between dry lab screening and wet lab research and development is driving commercialization out of Ontario to other regions with more appropriate lab space. Additionally, organizations must consider the cost of not addressing the gap. Manual data entry adds risk and can lead to rework, regulatory delays or failed tech transfers.
To successfully move between dry labs in one region and wet labs in another region, Canadian life sciences organizations need an informatics bridge and standardized data. A standardized informatics bridge ensures data integrity across the geographic divide. Adherence to ALCOA+ principles is crucial when Health Canada or the U.S. Food and Drug Administration asks exactly how a computational model arrived at a specific conclusion.
Geographic Specialization: Canada’s Wet West vs. Dry East
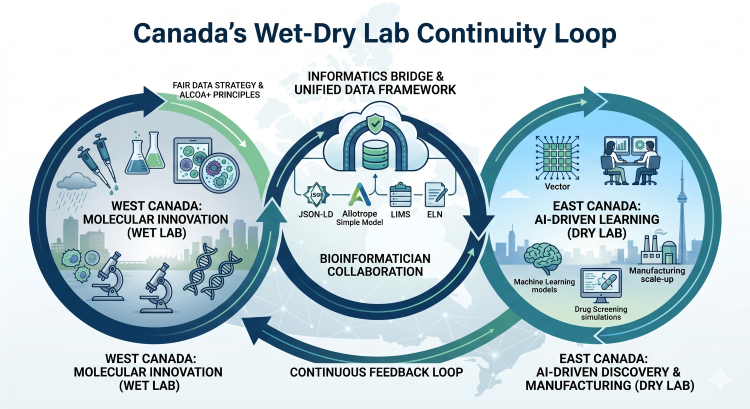
Vancouver’s weather is famously wet, and so is its current laboratory landscape. Much of the early-stage pharmaceutical research on drug targets in the greater Toronto area is currently conducted 3,000 kilometres away in Canada’s Pacific Northwest.
Vancouver has become a centre for molecular innovation. Companies like AbCellera, Amgen, and STEMCELL Technologies drive early-stage research, producing mass quantities of biological data. Similarly, back east, Montreal remains a stronghold of medicinal chemistry spearheaded by Pharmascience.
Conversely, the Greater Toronto Area hosts a concentration of computer scientists and medical physicists, supported by facilities such as the MaRS Centre. Life sciences organizations in the region are driving radioligand design (such as Fusion Pharmaceuticals and Alpha Cancer Technologies).
For investors and leaders, the message is clear: look West for discovery but look East for AI optimization. However, for organizations operating across these hubs, the distance isn't just physical—it's digital.
A unified informatics framework with a bioinformatician in the mix allows cross-functional teams to collaborate in real time and removes a bit of the black-box mystique. Bridging the gap with a cross-functional team helps the wet-lab scientists trust the AI model results and helps the dry-lab scientists be confident that their data can be experimentally verified.

Challenge: Breaking the Proprietary Data Cage
Beyond the shortage of wet lab space, there’s another hurdle to Canadian AI-driven discovery in life sciences. Data silos are intentionally created by instrument and system vendors’ proprietary data formats. Data in a vendor’s proprietary format can be invisible to ML models. Valuable wet lab research on one coast must be translated in some way so that it can be leveraged in a dry lab on the other coast. Edge computing and cloud-native data lakes are starting to address this hurdle, making access to the in silico model outputs faster on both coasts.
To remain competitive, Canadian organizations must adopt a vendor-neutral data strategy. Raw data must be extracted into standardized, vendor-agnostic formats such as the Allotrope Simple Model or JSON-LD. This lets a protein sequence or cellular assay generated in British Columbia and stored in a LIMS or ELN become AI ready once it hits a server in Ontario. Preserving data integrity during these transformations is paramount.
Harmonizing Discovery With Lab Informatics
The traditional approach to laboratory data management involves an ELN for discovery work and a LIMS for quality control. When these systems aren’t interfaced, data silos arise. Moving away from siloed ELN or LIMS data helps organizations advance toward a unified data framework. Harmonized, FAIR data bridges the gap between early-stage R&D data and clinical-stage requirements.

Modern laboratory informatics may use a middleware layer (often a scientific data platform) that harmonizes data across the entire lifecycle. This makes it possible to track a molecule from its first (dry) computational simulation to its first (wet) assay and eventually into clinical trials, without losing context.
Harmonizing your data ecosystem in this way has a few requirements:
- Standardized metadata schemas: These ensure that terms mean the same thing in every lab within an organization’s research pipeline. This is often accomplished by using an ontology.
- Automated data pipelines: These move data from the bench to the cloud automatically, reducing human error and system latency.
- Agile compliance systems: These satisfy Health Canada and U.S. Food and Drug Administration regulations without slowing down the fail fast mentality of R&D in the era of Lab 4.0.
Winning the Innovation Race
Bridging the divide between wet and dry labs is no longer just a matter of geography; it is a matter of digital infrastructure. The Canadian life sciences organizations that will dominate the next decade are those that stop treating wet and dry labs as separate silos and start treating them as a single, continuous data loop.
How could your organization make better use of your wet and dry lab data?







.png)


.png)

Comments